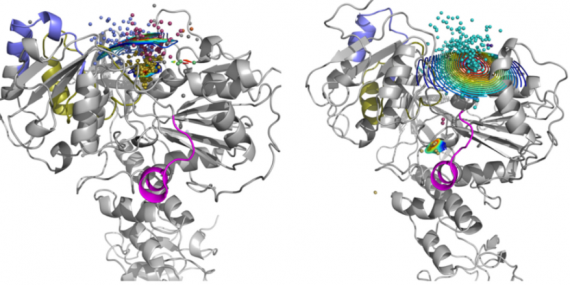
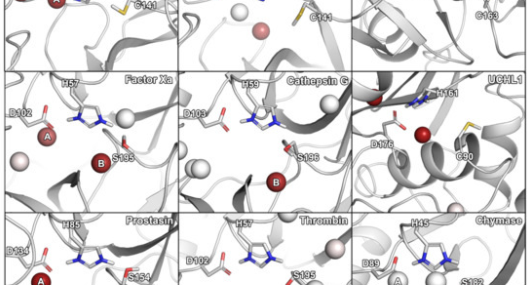
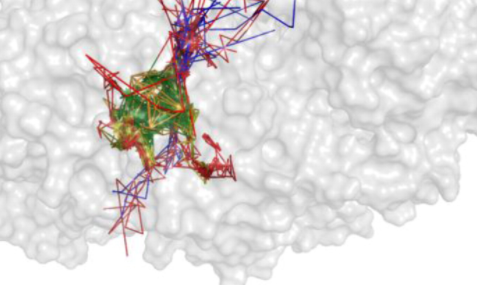
AQUA-DUCT Webinar: Tracking Solvent Pathways for Protein Engineering and Drug Design
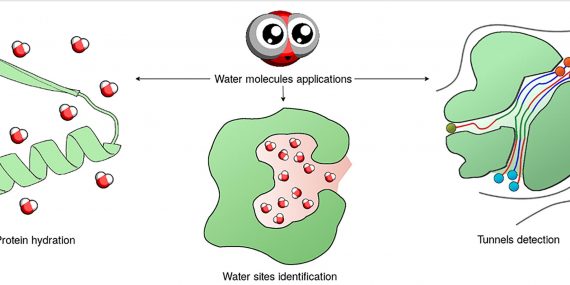
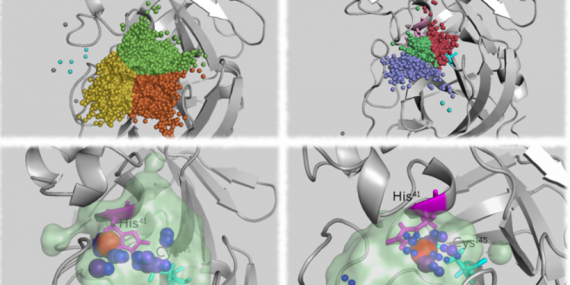
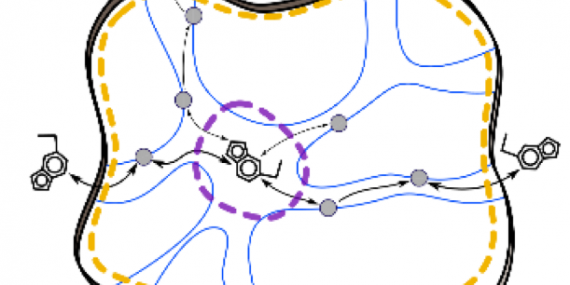
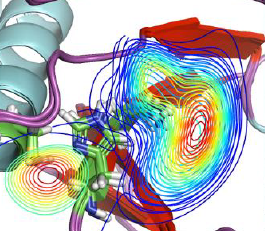
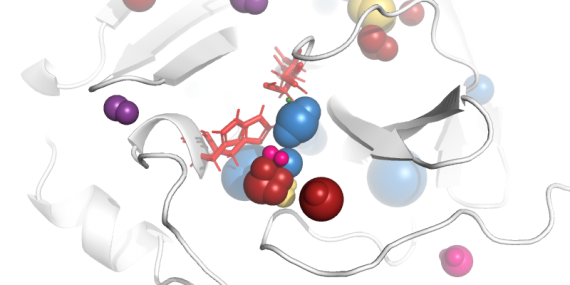
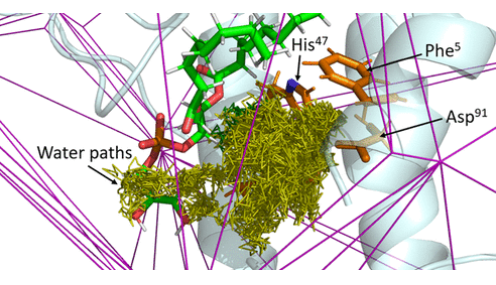
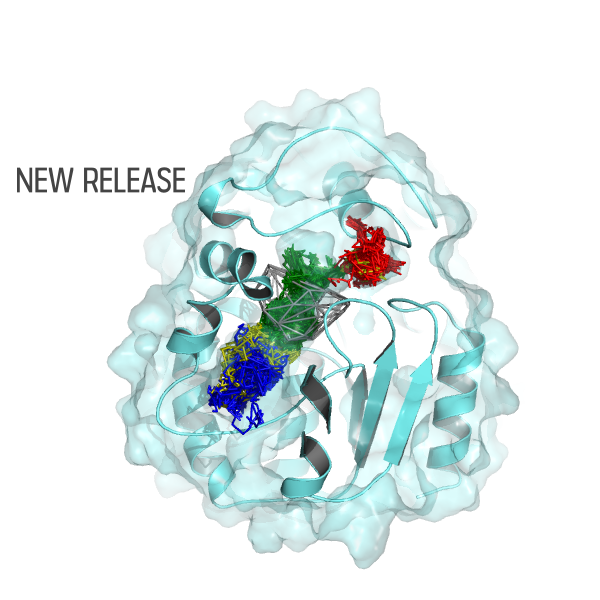
Join our upcoming webinar on 24 March 2026 at 15:00 CET to explore AQUA-DUCT! Workshop will introduce technical background and practical use of solvent tracking to investigate internal pathways, identify regions where small molecules become trapped and understand how water behavior supports enzymatic mechanism studies, protein engineering and drug design. More information and registration here