New citation
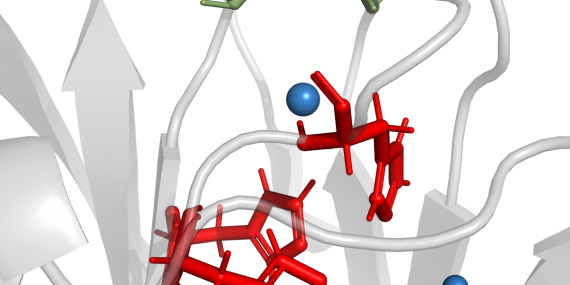
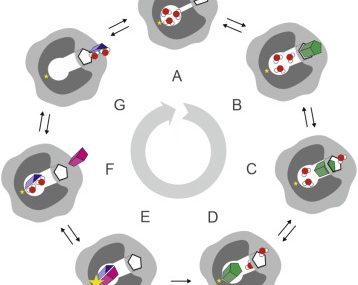
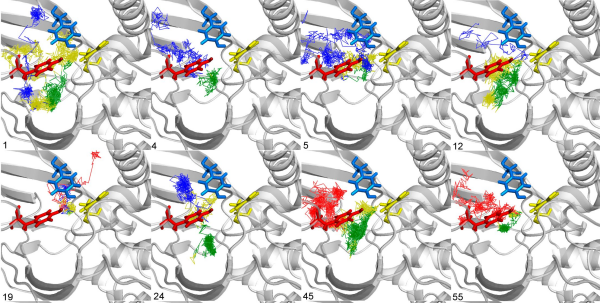
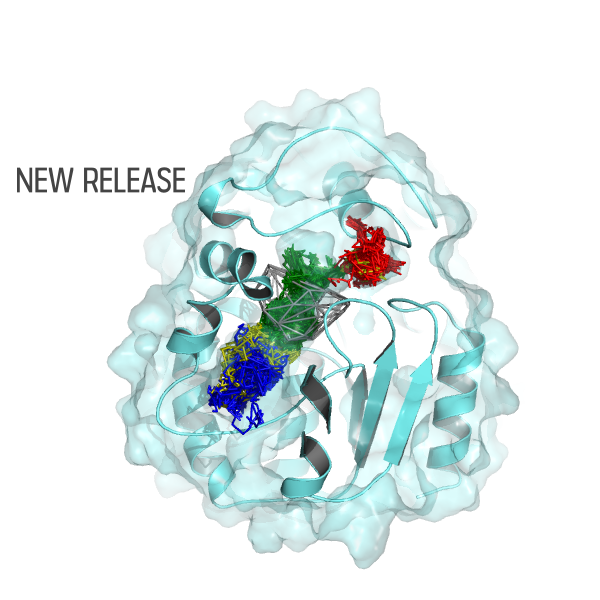
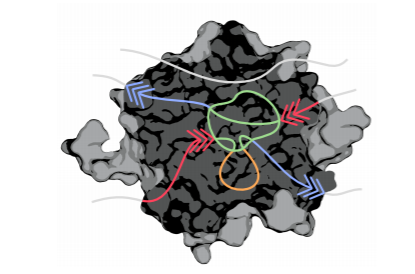
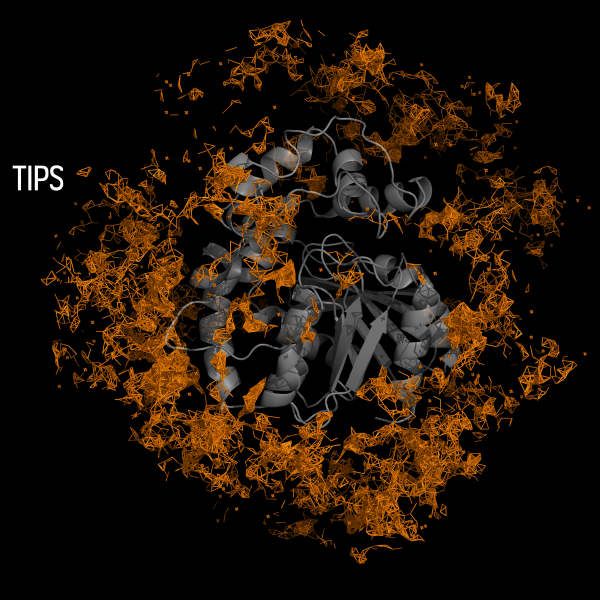
The AQUA-DUCT was used to study Hyperthermophilic Pyrococcus furiosus Phosphoglucose Isomerase. Recent paper was published in Biomolecules 2019, 9(6), 212. The cupin-type phosphoglucose isomerase (PfPGI) from the hyperthermophilic archaeon Pyrococcus furiosuscatalyzes the reversible isomerization of glucose-6-phosphate to fructose-6-phosphate. We investigated PfPGI using protein-engineering bioinformatics tools to select functionally-important residues based on correlated mutation analyses. A pair of amino acids in the periphery of…